
Comparison and clinical characteristics of COVID-19 between January and February 2020 in Wuhan, China
Introduction
A novel coronavirus disease [coronavirus disease 2019 (COVID-19)] outbreak has swept all over the world recently. It has been a huge threat to human life and health and has exerted enormous pressure on our socio-economic development. However, although the outbreak began in December 2019, it was well controlled in Wuhan, China, by the end of March. As of May 6, 2020, there were 50,333 cumulative confirmed cases of COVID-19, while, the death toll stood at 3,869 in Wuhan (1).
The severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) is a novel coronavirus. Until now, there have been two outbreaks caused by coronaviruses, i.e., severe acute respiratory syndrome (SARS) in 2002, and middle east respiratory syndrome (MERS) in 2012 (2). The SARS outbreak led to a reported about 8,098 cases, with 774 deaths, and a mortality rate of 9% (3). This comparted with MERS which led to about 2,519 infections and 866 deaths, and a mortality rate of about 34.4% (4). Previous studies have shown varying pathogenicity of both SARS and MERS over different time periods. Comparing the early stage (Nov 16, 2002–Jan 31, 2003), to the late stage of the outbreak (Jan 31, 2003–Feb 21, 2003), SARS-CoV showed reduced replicative ability and decreased virulence for host adaption in vitro, independent of the cell system examined (5,6). MERS-CoV was probably more sensitive to IFN-I (type I interferon) treatment at the later stage of the outbreak (7,8). In addition to coronaviruses, other viruses also exhibit varying pathogenicity over time, Ebola virus, belonging to a late-outbreak strain, exhibited reduced virulence and prolonged survival in experimental animals (9). H7N9, a novel avian influenza A virus, exhibited low pathogenicity in 2013, but was highly pathogenic in late 2016 showing altered virulence (10).
SARS-CoV-2 is a positive-sense, single-stranded RNA virus which has a high mutation rate similar to other RNA viruses. A great deal of genetic diversity has been found among clinical isolates (11). SARS-CoV-2 is able to encode four structural proteins, including spike (S), minor protein (E), matrix (M), and the nucleocapsid (N). Among them, the spike protein is responsible for entry into cells. Mutations have been frequently reported in the spike protein as well as variation in glycosylation sites (11-14). Whether persistent mutations of SARS-CoV-2 will lead to changes in the transmission, the symptoms, or the severity of the disease is not yet clear.
After December 2019, strict measures were taken to prevent crowd gathering, and self-segregation was accepted, to curb the spread of SARS-CoV-2 in Wuhan city. The influx of medical teams provided support for hospital care. These efforts ultimately resulted in control of the outbreak. On April 4, 2020, no new cases were diagnosed with the COVID-19 in Wuhan city (15). Previous studies have shown that the SARS-CoV and MERS-CoV both demonstrated reduced pathogenicity over time. However, it was unclear whether the pathogenicity of SARS-CoV-2 would also alter over time. Therefore, we aimed to compare clinical characteristics and outcome of COVID-19 patients over 2 different time periods in order to detect any change in pathogenicity of SARS-CoV-2. COVID-19 was an acute and serious assault on humanity inflicting enormous loss of life but also heavy economic and social burdens. Furthermore, it is not known how long it will last. Therefore, understanding any changes in pathogenicity and transmission of SARS-CoV-2 would be important for adjustment of preventative measures and to minimize any waste of medical resource. It is likely that any change of clinical characteristics and outcomes, if corroborated with change in viral pathogenicity with emerging mutations, would be valuable in predicting future disease progression.
We present the following article in accordance with the STROBE reporting checklist (available at http://dx.doi.org/10.21037/apm-20-2222).
Methods
Study population
We recruited 1,347 patients who were admitted to the Renmin Hospital of Wuhan University between 30th January, 2020 and 16st March, 2020: all patients being discharged before 31st March, 2020. However, we excluded patients as follows: 167 patients who were transferred to Fangcang shelter hospital around 9th February (because of missing outcome data), 156 patients who were transferred from Fangcang shelter hospital in early March because of missing initial data, 44 patients with repeatedly negative pharyngeal swab SARS-CoV-2 test results, and 84 patients because of other essential missing data. Therefore, we finally considered 896 patients diagnosed with COVID-19 (Figure 1). The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Medical Ethics Committee of Renmin Hospital of Wuhan University, Wuhan, China (No. WDRY2020-K019) and individual consent for this retrospective analysis was waived.
Procedures
Trained physicians and medical students reviewed the demographics, clinical characteristics, radiologic and laboratory findings, treatments administered, and clinical outcomes from electronic medical records. Symptoms appearing before admission were also recorded. Vital signs and disease severity status were recorded during admission. The results of blood tests and radiologic findings, following admission, were analyzed. The highest level of oxygen support during hospitalization was recorded. During the whole study period, patients’ confidentiality was safeguarded. Nasopharyngeal swab samples were collected from patients on or before admission and during their hospital stay, and repeated as required. The double nucleic acid detection kit (Bio-Germ, Shanghai, China) was used to detect SARS-CoV-2 using the open reading frame 1ab (ORF1ab)/and nucleocapsid protein (N) genes. Nearly all patients underwent routine blood tests, including full blood count, coagulation tests, renal and liver biochemical profile, C-reactive protein (CRP), D-dimers, CD3 and CD4 cell counts, procalcitonin (PCT), and radiology including chest computed tomography.
Definitions
COVID-19 was diagnosed based on a positive result of nasopharyngeal swab samples at any time during the clinical course plus one of several criteria including: fever, respiratory symptoms and bilateral or multiple ground-glass opacities on chest CT scan. The severity of COVID-19 was defined based on the Chinese National COVID-19 Diagnosis and Treatment Guidelines version-7 (16). Mild disease patients showed only mild symptoms. Moderate disease patients presented with only slight symptoms but an abnormal computed tomography (CT) scan. Severe disease patients exhibited dyspnea, a respiratory rate ≥30 times/min, SpO2 (percutaneous oxygen saturation) ≤93%, and an oxygenation index (PaO2/FiO2) ≤300 mmHg. Critical illness patients presented with respiratory failure, shock or multiple organ failure ultimately requiring mechanical ventilation and intensive care. Fever was defined as a temperature equal or greater than 37.3 °C. Acute respiratory distress syndrome (ARDS) and acute kidney injury (AKI) were defined according to the Berlin Definition (17) and the Kidney Disease: Improving Global Outcomes (KDIGO) clinical practice guidelines (18), respectively. Sepsis was defined according to 2016 Third International Consensus Definition (19). Acute heart failure (AHF) and acute liver failure (ALF) were defined based on Recommendations on pre-hospital and early hospital management of AHF (20) and the American Gastroenterological Association Institute Guidelines for the Diagnosis and Management of Acute Liver Failure (21), respectively. Hypoalbuminemia was defined as a blood albumin less than 35 g/L. Shock was defined as systolic blood pressure lower than 90 mmHg or mean arterial pressure lower than 65 mmHg.
Statistical analysis
Categorical data such as sex, symptoms, comorbidity, vital signs, disease severity status, abnormal blood test results, imaging features, treatments given and outcomes were compared between patients with onset of illness in January and patients with onset of illness in February 2020 using the χ2 test. Non-normal distributed, continuous data, including age, time from illness onset to hospital admission, blood test results, time from illness onset to discharge or death, were compared using the Mann-Whitney-Wilcoxon test. Categorical variables were represented as frequencies and percentages, and abnormally distributed continuous variables were described by the median and interquartile range (IQR). Survival data were analyzed by the Kaplan-Meier plot. Univariate and multivariate logistic regressions were performed to evaluate if the time of symptoms onset was an independent risk factor for in-hospital death. Based on previous findings, and clinical experience, age (22), sex, presence of comorbidity, and time from illness onset to hospital admission were chosen for univariate and multivariate analysis to eliminate confounders. The factors with statistical significance in the univariate analysis were included in the multivariate analysis. All statistical data were analyzed using SPSS (v23) software and figures plotted using GRAPHPAD PRISM (v8.0). P values less than 0.05 were considered as statistically significant.
Results
Demographics and clinical characteristics on admission
A total of 896 patients with a confirmed diagnosis of COVID-19 were enrolled. The median age was 60 (IQR, 47–69) years, 431 (48.1%) were male. The most common symptoms were fever (80.4%) and cough (50.4%). Some patients showed symptoms related to the digestive tract, including diarrhea (17.7%), nausea or vomiting (5.8%), abdominal pain and distention (2.3%) and lack of appetite (13.1%). A few patients had symptoms such as itchiness of the eyes (0.8%) (Table 1).
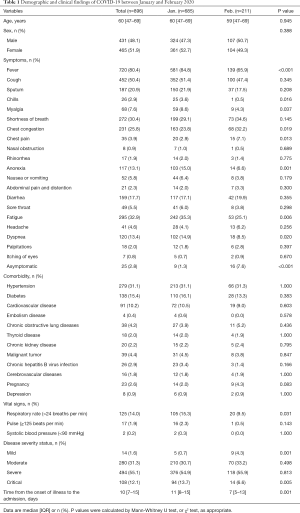
Full table
According to time of illness onset, the 896 patients were divided into group A (n=685, 76.5%; patients with onset of illness in January 2020) and group B (n=211, 23.5%; patients with onset of illness in February 2020). The median ages were 60 (IQR, 47–69) years and 59 (IQR, 47–69) years respectively. The ratio of males and females was approximately 1:1 in both group A and group B. Compared with group B, group A had a higher incidence of fever (84.8% vs. 65.9%, P<0.001), chills (3.6% vs. 0.5%, P=0.016), myalgia (8.6% vs. 4.3%, P=0.037) but a lower rate of chest congestion (23.8% vs. 32.2%, P=0.019) and chest pain (2.9% vs. 7.1%, P=0.013). Nonspecific symptoms, including loss of appetite (15.0% vs. 6.6%, P=0.001) and fatigue (35.3% vs. 25.1%, P=0.006), were significantly less common in patients in group B. Dyspnea was significantly less common (14.9% vs. 8.5%, P=0.020), and asymptomatic patients were significantly more common (1.3% vs. 7.6%, P<0.001) in group B (Table 1).
Among all 896 COVID-19 patients, 470 (52.5%) had comorbidities. Hypertension (31.1%), diabetes (15.4%) and cardiovascular disease (10.2%) were the most common comorbidities. However, there was no significant difference in comorbidities between two groups (Table 1).
Compared with group B, group A had a higher rate of mortality (10.2% vs. 5.2%, P=0.028), a higher rate of critical illness (13.7% vs. 6.6%, P=0.005), but a lower rate of patients with mild disease (0.7% vs. 4.3%, P=0.001). The percentages of moderate patients and severe patients did not differ significantly between the two groups. There was a significant difference in time from the onset of illness to the admission (P=0.001); with a median of 11 (IQR, 8–15) days in group A and 7 (IQR, 5–13) days in group B. The percentage of patients with a respiratory rate higher than 24 breaths per min, during admission, was increased significantly in group A (15.3% vs. 9.5%, P=0.031) (Table 1).
Laboratory findings and image feature
Laboratory results were collected during the admission. Considering all the patients, the most common abnormality was an increase in D-dimer (63.3%), about half of patients demonstrated an elevated CRP (60%), increased fibrinogen (53.6%), raised lactate dehydrogenase (LDH) (50.1%) and decreased lymphocyte count (48.7%). However, relatively few patients showed kidney injury with an increased level of creatinine (7.7%) or urea (13.6%) (Table 2).
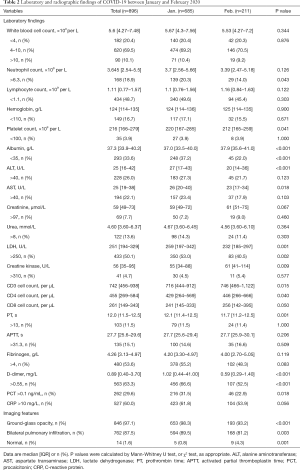
Full table
Compared with group B, group A patients had a higher percentage of increased D-dimer (66.6% vs. 52.5%, P<0.001) and PCT (31.5% vs. 22.9%, P=0.018), as well as neutrophilia (20.3% vs. 14.0%, P=0.043) and elevated levels of LDH (53.0% vs. 40.5%, P=0.002). Group B patients had higher CD3 (P=0.015) and CD4 cell counts (P=0.040) than group A patients. Liver injury indices [in terms of mean alanine aminotransferase (ALT) and aspartate transaminase (AST) values rather than the number of patients with abnormally high levels], including ALT (P<0.001) and AST (P=0.018), were significantly higher in group A than in group B. More hypoalbuminemia occurred in group A patients (37.2% vs. 22.0%, P<0.001). However, there was no significantly difference between the two groups as regards total white blood cell count, lymphocyte count and CRP (Table 2).
Compared with group B, more CT scans in group A showed bilateral pulmonary infiltrates (89.5% vs. 81.2%, P=0.003) and ground-glass opacities (98.3% vs. 93.2%, P<0.001), while group B patients more frequently had normal CT scans (0.8% vs. 4.3%, P=0.001) (Table 2).
Treatments and outcomes
Of the 896 patients, most received antibiotics (76.9%), Lianhua Qingwen (a traditional Chinese medicine which contains 11 herbs and two mineral Chinese medicine) capsules (71.8%) and antiviral treatment (93.4%) which included oseltamivir, Arbidol and ganciclovir. Compared to group B, group A received more treatment with antibiotics (79.1% vs. 69.7%, P=0.009), corticosteroids (41.3% vs. 33.2%, P=0.036), intravenous immunoglobin (48.9% vs. 28.9%, P<0.001) and interferon α (18.7% vs. 10.9%, P=0.008). We recorded the highest level of oxygen support, and 701 (78.2%) patients required supplemental oxygen therapy. More high-flow nasal cannula oxygen therapy (10.4% vs. 5.2%, P=0.028) and more non-invasive mechanical ventilation (4.1% vs. 0.9%, P=0.027) were administered in group A patients (Table 3).
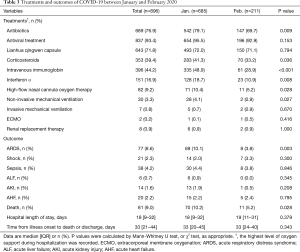
Full table
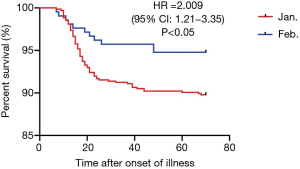
Among all the patients, median hospital stay was 18 (IQR, 9–32) days, while 81 patients (9.0%) died. Seventy-seven (8.6%) patients developed ARDS, sepsis occurred in 38 (4.2%) and shock was observed in 21 (2.3%) patients. Among the patients with shock, this was due to sepsis in 9 (42.9%), related to heart failure in 10 (47.6%) and caused by acute gastrointestinal bleeding in 3 (14.3%). None of the patients suffering shock survived. The higher rate of mortality in group A compared with group B patients (10.2% vs. 5.2%, P=0.028) is shown in Figure 2. Group A patients had a higher incidence of developing ARDS (10.1% vs. 3.8%, P=0.003). Considering all the patients, median time from illness onset to death or discharge was 33 (IQR, 21–44) days and there was no significant difference between the two groups (Table 3).
Risk factors associated with in-hospital death
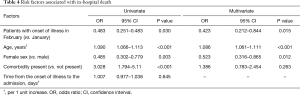
In a univariate analysis, higher odds of in-hospital death were associated with older age (odds ratio 1.09, 95% CI: 1.066–1.113, per year increase; P<0.001) and presence of comorbidity (odds ratio 3.028, 95% CI: 1.794–5.11; P<0.001). Female sex (odds ratio 0.485, 95% CI: 0.302–0.779; P=0.003) and group B timing (odds ratio 0.483, 95% CI: 0.251–0.483; P=0.03) were associated with decreased risk of in-hospital death. Time from illness onset to hospital admission was not a risk factor for in-hospital death. In multivariate analysis, we found that older age (odds ratio 1.086, 95% CI: 1.061–1.111, per year increase; P<0.001) was associated with higher odds of in-hospital death, female sex (odds ratio 0.523, 95% CI: 0.316–0.865; P=0.012) and group B timing (odds ratio 0.423, 95% CI: 0.212–0.844; P=0.015) gave lower odds of in-hospital death (Table 4).
Full table
Discussion
In our study, we found a significant difference in severity of COVID-19 in patients diagnosed during the 2 different time periods. Comparing patients with onset of illness in January, those with onset of illness in February 2020 had less severe disease; with a lower mortality, a lower rate of critical illness, lower blood levels of neutrophils, D-dimer, LDH, PCT, ALT, AST and higher CD3 and CD4 cell counts. In addition, we observed a significant increase in mild disease, in asymptomatic patients and in patients with normal CT chest scans in those whose illness began in February 2020. We also found that older age, male sex, and illness onset in January 2020, were independent factors for in-hospital death.
Increased neutrophil count was positively related with poor outcome in COVID-19. The previous study by Wu et al. demonstrated that neutrophilia is a risk factor for progression to ARDS and death from ARDS in COVID-19 patients (23). Neutrophilia was also a risk factor for intubation in SARS patients (24). Severity of lung damage correlated positively with peripheral blood neutrophil count in patients with MERS (25,26). Neutrophils produce chemokines and cytokines which combat coronaviruses but at the same time can contribute to an excess inflammatory response resulting in multiple organ injury. As previously shown, an elevated D-dimer was an independent risk factor for mortality in COVID-19 patients (23,27). Coagulation disorders are prominent in COVID-19 patients (28) and non-survivors had higher levels of D-dimers, FDP, and a prolonged prothrombin time (PT) and activated partial thromboplastin time (APTT) (29). Possible mechanisms include viral triggering of the inflammatory cascade and secondary infection causing microvascular injury and release of procoagulant factors. Evidence suggested that higher CD3 and CD4 T-cell counts protected patients from developing ARDS (23). A recent electron microscopy study localized SARS-CoV-2 to CD4 cells in COVID-19 patients, however, there was no viral replication (30) suggesting that CD4 cells might control SARS-CoV-2 replication. The incidence of secondary liver damage was positively related to poor outcome in COVID-19 patients (31). A recent study showed that liver damage was caused by activated cytotoxic T lymphocytes triggered by SARS-CoV-2 (32), in addition to drug-induced liver injury. Increased LDH was also associated with poor outcome in COVID-19 patients (23,27). Mild patients and asymptomatic patients were more common in the group with onset of illness in February 2020. A recent study of patients with mild or asymptomatic COVID-19, showed virus predominantly in the upper respiratory tract (33,34). Other studies have shown a worse prognosis in patients over the age of 60 years, consistent with our study (22). Males with COVID-19 have worse outcomes and higher mortality in many studies (35-37) possibly due to differences in chromosomes, reproductive organs, or the related sex steroids (38).
Mortality, symptoms and abnormal results of laboratory tests were all reduced in patients with onset of illness in February compared with those with onset of illness in January 2020. We speculate that the improved outcome might relate to reduced SARS-CoV-2 virulence. While one month may seem like a very short-time frame for altered viral virulence there is considerable evidence that significant changes in pathogenicity can occur over periods as short as 3 months (39).
Recently, variants (40), mutations (41) and genetic diversity (11,42) have increasingly been described in SARS-CoV-2, which could explain diminishing virulence. A 382-nt deletion (43) and a 29-nt deletion both covering open reading frame 8 (ORF8) have appeared during SARS-CoV-2 and SARS-CoV evolution respectively. The 29-nt deletion resulted in decreased replicative ability of SARS-CoV in-vitro, independent of the cell system (6). The 382-nt deletion may also reduce virulence of SARS-CoV-2. An 81-nucleotide deletion in ORF7 causing a 27 amino-acid in-frame deletion (44), may also change SARS-CoV-2 pathogenicity. In the United States, researchers found a discrepancy in mortality of COVID-19 patients between the East Coast and West Coast of America. They speculated that this was probably due a change in the viral spike protein (45). Yao et al. similarly found greatly altered pathogenicity of 11 SARS-CoV-2 viral isolates, infecting Vero-E6 cells in vitro (46). In addition, reduced virulence likely affected antibody generation. Improved clinical outcome in COVID-19 patients treated with convalescent plasma has been well documented (47).
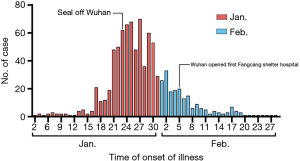
In addition to diminishing virulence, other factors may have caused a difference in severity of COVID-19 over these different time periods. At an early stage of the outbreak, the time from illness onset to hospital admission was longer, probably because of a shortage of medical resources and lack of awareness of COVID-19 which may have led to more severe illness. Moreover, in the later stage of the outbreak, more strict management was employed to curb the spread of infection (48), with more Fangfang shelter hospitals established and improved healthcare so cases may have been detected earlier. However, our hospital was designated to accept COVID-19 patients with severe and critical disease. Furthermore, accumulated clinical experience and published research led to improved outcomes (Figure 3).
We acknowledge a few limitations of our study. Firstly, in the early stages of the outbreak, many patients who showed clinical improvement were transferred to other isolation sites for further observation; thus, we did not obtain the time of viral clearance. Secondly, due to the retrospective nature of the study, some data are missing from analysis. Thirdly, some patients with onset of illness in February were not included as they remain in hospital. Furthermore, our hospital preferentially accepted more severe and critical illness COVID-19 patients in the later stage of outbreak, therefore the spectrum of disease severity and mortality in our study did not reflect the actual situation in Wuhan city. Although our study had a limited sample size, it was conducted at a single-center.
Conclusions
We analyzed the clinical characteristics and outcomes of COVID-19 during different time periods in Wuhan. We found that patients with onset of illness in February showed improved clinical outcome and decreased risk for in-hospital death compared to patients with onset of illness in January 2020. This would be consistent with reduced viral virulence although other factors need to be taken into consideration. Further studies, with viral sequencing and population immunity surveillance are required to confirm our hypothesis.
Acknowledgments
We would like to express our gratitude to EditSprings (
Funding: This work was supported by the National Key Research and Development Plan for the Emergency Management of Novel Coronavirus Pneumonia (No. 2020YFC0845100)
Footnote
Reporting Checklist: The authors have completed the STROBE reporting checklist. Available at http://dx.doi.org/10.21037/apm-20-2222
Data Sharing Statement: Available at http://dx.doi.org/10.21037/apm-20-2222
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/apm-20-2222). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Medical Ethics Committee of Renmin Hospital of Wuhan University, Wuhan, China (No. WDRY2020-K019) and individual consent for this retrospective analysis was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Available online: http://wjw.hubei.gov.cn/fbjd/dtyw/202005/t20200507_2260769.shtml. Accessed May 6 2020.
- Zaki AM, van Boheemen S, Bestebroer TM, et al. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med 2012;367:1814-20. [Crossref] [PubMed]
- Fehr AR, Perlman S. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol 2015;1282:1-23. [Crossref] [PubMed]
- Rabaan AA, Al-Ahmed SH, Haque S, et al. SARS-CoV-2, SARS-CoV, and MERS-COV: A comparative overview. Infez Med 2020;28:174-84. [PubMed]
- Chinese SARS Molecular Epidemiology Consortium. Molecular evolution of the SARS coronavirus during the course of the SARS epidemic in China. Science 2004;303:1666-9. [Crossref] [PubMed]
- Muth D, Corman VM, Roth H, et al. Attenuation of replication by a 29 nucleotide deletion in SARS-coronavirus acquired during the early stages of human-to-human transmission. Sci Rep 2018;8:15177. [Crossref] [PubMed]
- Niemeyer D, Zillinger T, Muth D, et al. Middle East respiratory syndrome coronavirus accessory protein 4a is a type I interferon antagonist. J Virol 2013;87:12489-95. [Crossref] [PubMed]
- Payne DC, Biggs HM, Al-Abdallat MM, et al. Multihospital Outbreak of a Middle East Respiratory Syndrome Coronavirus Deletion Variant, Jordan: A Molecular, Serologic, and Epidemiologic Investigation. Open Forum Infect Dis 2018;5:ofy095 [Crossref] [PubMed]
- Marzi A, Chadinah S, Haddock E, et al. Recently Identified Mutations in the Ebola Virus-Makona Genome Do Not Alter Pathogenicity in Animal Models. Cell Rep 2018;23:1806-16. [Crossref] [PubMed]
- Bisset AT, Hoyne GF. Evolution and Adaptation of the Avian H7N9 Virus into the Human Host. Microorganisms 2020;8:778. [Crossref] [PubMed]
- Sheikh JA, Singh J, Singh H, et al. Emerging genetic diversity among clinical isolates of SARS-CoV-2: Lessons for today. Infect Genet Evol 2020;84:104330 [Crossref] [PubMed]
- Li Q, Wu J, Nie J, et al. The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity. Cell 2020;182:1284-1294.e9. [Crossref] [PubMed]
- Dawood AA. Mutated COVID-19 may foretell a great risk for mankind in the future. New Microbes New Infect 2020;35:100673 [Crossref] [PubMed]
- Saha P, Banerjee AK, Tripathi PP, et al. A virus that has gone viral: amino acid mutation in S protein of Indian isolate of Coronavirus COVID-19 might impact receptor binding, and thus, infectivity. Biosci Rep 2020;40:BSR20201312 [Crossref] [PubMed]
- Available online: https://baijiahao.baidu.com/s?id=1664839948318758995&wfr=spider&for=pc. Accessed May 1 2020.
- Chinese management guideline for COVID-19 (version 7.0). Available online: http://www.nhc.gov.cn/yzygj/s7653p/202003/46c9294a7dfe4cef80dc7f5912eb1989.shtml
- ARDS Definition Task Force. Acute respiratory distress syndrome: the Berlin Definition. JAMA 2012;307:2526-33. [PubMed]
- Khwaja A. KDIGO clinical practice guidelines for acute kidney injury. Nephron Clin Pract 2012;120:c179-84. [PubMed]
- Seymour CW, Liu VX, Iwashyna TJ, et al. Assessment of Clinical Criteria for Sepsis: For the Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 2016;315:762-74. [Crossref] [PubMed]
- Mebazaa A, Yilmaz MB, Levy P, et al. Recommendations on pre-hospital and early hospital management of acute heart failure: a consensus paper from the Heart Failure Association of the European Society of Cardiology, the European Society of Emergency Medicine and the Society of Academic Emergency Medicine--short version. Eur Heart J 2015;36:1958-66. [Crossref] [PubMed]
- Flamm SL, Yang YX, Singh S, et al. American Gastroenterological Association Institute Guidelines for the Diagnosis and Management of Acute Liver Failure. Gastroenterology 2017;152:644-7. [Crossref] [PubMed]
- Liu Y, Mao B, Liang S, et al. Association between age and clinical characteristics and outcomes of COVID-19. Eur Respir J 2020;55:2001112 [Crossref] [PubMed]
- Wu C, Chen X, Cai Y, et al. Risk Factors Associated with Acute Respiratory Distress Syndrome and Death in Patients With Coronavirus Disease 2019 Pneumonia in Wuhan, China. JAMA Intern Med 2020;180:934-43. Erratum in: JAMA Intern Med 2020 Jul 1;180(7):1031. doi: 10.1001/jamainternmed.2020.1429. [Crossref] [PubMed]
- Wang YH, Lin AS, Chao TY, et al. A cluster of patients with severe acute respiratory syndrome in a chest ward in southern Taiwan. Intensive Care Med 2004;30:1228-31. [Crossref] [PubMed]
- Min CK, Cheon S, Ha NY, et al. Comparative and kinetic analysis of viral shedding and immunological responses in MERS patients representing a broad spectrum of disease severity. Sci Rep 2016;6:25359. [Crossref] [PubMed]
- Kim ES, Choe PG, Park WB, et al. Clinical Progression and Cytokine Profiles of Middle East Respiratory Syndrome Coronavirus Infection. J Korean Med Sci 2016;31:1717-25. [Crossref] [PubMed]
- Zhou F, Yu T, Du R, et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 2020;395:1054-62. [Crossref] [PubMed]
- Zhang Y, Xiao M, Zhang S, et al. Coagulopathy and Antiphospholipid Antibodies in Patients with Covid-19. N Engl J Med 2020;382:e38 [Crossref] [PubMed]
- Tang N, Li D, Wang X, et al. Abnormal coagulation parameters are associated with poor prognosis in patients with novel coronavirus pneumonia. J Thromb Haemost 2020;18:844-7. [Crossref] [PubMed]
- Banerjee A, Nasir JA, Budylowski P, et al. Isolation, Sequence, Infectivity, and Replication Kinetics of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg Infect Dis 2020;26:2054-63. [Crossref] [PubMed]
- Chen N, Zhou M, Dong X, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 2020;395:507-13. [Crossref] [PubMed]
- Bangash MN, Patel J, Parekh D. COVID-19 and the liver: little cause for concern. Lancet Gastroenterol Hepatol 2020;5:529-30. [Crossref] [PubMed]
- Wölfel R, Corman VM, Guggemos W, et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020;581:465-9. Erratum in: Nature 2020 Dec;588(7839):E35. doi: 10.1038/s41586-020-2984-3. [Crossref] [PubMed]
- Qiu J. Covert coronavirus infections could be seeding new outbreaks. Nature 2020; Epub ahead of print. [Crossref] [PubMed]
- Jin JM, Bai P, He W, et al. Gender Differences in Patients With COVID-19: Focus on Severity and Mortality. Front Public Health 2020;8:152. [Crossref] [PubMed]
- Meng Y, Wu P, Lu W, et al. Sex-specific clinical characteristics and prognosis of coronavirus disease-19 infection in Wuhan, China: A retrospective study of 168 severe patients. PLoS Pathog 2020;16:e1008520 [Crossref] [PubMed]
- Qin L, Li X, Shi J, et al. Gendered effects on inflammation reaction and outcome of COVID-19 patients in Wuhan. J Med Virol 2020;92:2684-92. [Crossref] [PubMed]
- Gebhard C, Regitz-Zagrosek V, Neuhauser HK, et al. Impact of sex and gender on COVID-19 outcomes in Europe. Biol Sex Differ 2020;11:29. [Crossref] [PubMed]
- Korber B, Fischer WM, Gnanakaran S, et al. Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell 2020;182:812-827.e19. [Crossref] [PubMed]
- Forster P, Forster L, Renfrew C, et al. Phylogenetic network analysis of SARS-CoV-2 genomes. Proc Natl Acad Sci U S A 2020;117:9241-3. [Crossref] [PubMed]
- Benvenuto D, Angeletti S, Giovanetti M, et al. Evolutionary analysis of SARS-CoV-2: how mutation of Non-Structural Protein 6 (NSP6) could affect viral autophagy. J Infect 2020;81:e24-7. [Crossref] [PubMed]
- Shen Z, Xiao Y, Kang L, et al. Genomic Diversity of Severe Acute Respiratory Syndrome-Coronavirus 2 in Patients with Coronavirus Disease 2019. Clin Infect Dis 2020;71:713-20. [Crossref] [PubMed]
- Su YC, Anderson DE, Young BE, et al. Discovery of a 382-nt deletion during the early evolution of SARS-CoV-2. bioRxiv 2020. doi: https://doi.org/
10.1101/2020.03.11.987222 . - Holland LA, Kaelin EA, Maqsood R, et al. An 81-Nucleotide Deletion in SARS-CoV-2 ORF7a Identified from Sentinel Surveillance in Arizona (January to March 2020). J Virol 2020;94:e00711-20. [Crossref] [PubMed]
- Brufsky A. Distinct viral clades of SARS-CoV-2: Implications for modeling of viral spread. J Med Virol 2020;92:1386-90. [Crossref] [PubMed]
- Yao H, Lu X, Chen Q, et al. Patient-derived mutations impact pathogenicity of SARS-CoV-2. medRxiv, 2020. doi:
10.1101/2020.04.14.20060160 . - Shen C, Wang Z, Zhao F, et al. Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA 2020;323:1582-9. [Crossref] [PubMed]
- Tian H, Liu Y, Li Y, et al. An investigation of transmission control measures during the first 50 days of the COVID-19 epidemic in China. Science 2020;368:638-42. [Crossref] [PubMed]



