
Tigecycline in combination with other antibiotics against clinical isolates of carbapenem-resistant Klebsiella pneumoniae in vitro
Introduction
Klebsiella pneumoniae (KP) are Enterobacteriaceae responsible for various human infections such as bacteremia, meningitis, and pneumonia (1). Standard treatment involves the use of carbapenems whose wide application has led to the emergence of carbapenem-resistant Enterobacteriaceae (CRE) (2). The resistance occurring in China is mainly due to the emergence of carbapenems hydrolases producing Klebsiella pneumoniae carbapenems (KPC), for example. The infection caused by carbapenem-resistant Klebsiella pneumoniae (CRKP) is complicated. CRKP usually carries multiple drug resistance genes which limit the treatment effect. At present, only a few drugs, such as colistin (COL), tigecycline (TGC), some aminoglycosides, and ceftazidime-avibactam, still have favorable in vitro activity against CPKP. The potential nephrotoxicity of COL and aminoglycosides restricts their broad use, especially among elderly patients with renal insufficiency. Ceftazidime–avibactam only inhibits the activity of class A enzymes. TGC, the 9-t-butylglycylamido derivative of minocycline, is regarded as the last resort for the management of difficult-to-treat carbapenem-resistant isolates infections (3). This limitation in treatment options further amplifies the need for new antibiotics (4,5). The dearth of novel antibiotics introduced to the market has been attributed to regulatory hurdles, high research costs, and low investment returns (6). A possible solution is drug repurposing. Since an approved drug has been cleared in terms of safety, pharmacological profile, and manufacturing process, it can be rapidly made available for a new disease indication (7). Based on this, we screened TGC combinations with other antibiotics, such as COL, meropenem (MEM), amikacin (AMK), and levofloxacin (LEV), to study antibacterial activity against clinical isolates of CRKP in vitro. Our research may provide laboratory data on antibiotic the susceptibility for drug-resistant patients and compensate for the lack of domestic antibiotic-related studies in the east of Heilongjiang.
Methods
Bacterial isolates
Forty CRKP isolates were collected from patients at the 1980-bed First Affiliated Hospital of Jiamusi University in Heilongjiang Province, northeast China, from October 2015 to January 2019. The isolates were identified as CRKP isolates by the Vitek 2 system (bioMérieux) and the AST-GN card (bioMérieux, France). Carbapenemase resistance (either to ertapenem, imipenem, or meropenem) was determined according to Clinical and Laboratory Standard Institutes (CLSI-2016) criteria.
Antimicrobial susceptibility testing
The concentration ranges for the antibiotics tested were 0.25–128 µg/mL for TGC, 0.0625–64 µg/mL for COL, 0.5–256 µg/mL for MEM, 0.03125–16,384 µg/mL for AMK, and 0.25–128 µg/mL for LEV. The minimum inhibitory concentrations (MICs), defined as the lowest compound concentrations (µg/mL) required to stop bacterial growth, of the five antibiotics tested were determined using the microbroth dilution method per CLSI recommendations (8). Strain KP ATCC 700603 was ESBL-positive and was used as the reference strain. The MEM, AMK, and LEV results were interpreted based on CLSI criteria (8), whereas the TGC and COL results were interpreted based on the European Committee on Antimicrobial Susceptibility Testing (EUCAST) breakpoint recommendations (9). Finally, the MIC results were read on an enzyme-labeled instrument via optical density (OD) at 570 nm.
Checkerboard technique
TGC was tested for synergistic activity against 10 CRKP isolates via the checkerboard technique with the TGC-COL, TGC-MEM, TGC-AMK, and TGC-LEV combinations. The concentration ranges were based on the MICs determined above. Fifty microliters of each antibiotic at 5 increasing (4-fold) concentrations (0.125× MIC to 2× MIC) were used, and each well was inoculated with 100 µL of a 7.5×105 CFU/mL suspension of the test CRKP isolates in a final volume of 200 µL in duplicate. Results were measured using an enzyme-labeled instrument through OD at 570 nm after incubating at 37 °C for 24 hours.
The effects of the different antimicrobial combinations were defined according to the fractional inhibitory concentration index (FICI) as follows: FICI ≤0.5, synergism; 0.5< FICI ≤1, additive; 1< FICI ≤2, indifferent; or FICI >2, antagonistic (10).
Antibacterial time-kill assay
The antibacterial time-kill assay was used to assess the results of the antibiotic monotherapies and TGC combinations against 4 CRKP isolates (including 3 KPC-producing isolates and 1 KPC-free isolate) based on CLSI guidelines. Antibiotic concentrations were calculated using the mean value of the steady-state concentrations of the nonprotein-bound drug in humans as described previously (11). The monotherapies were carried out with TGC 0.1 and 1 mg/L; COL 0.25 and 1 mg/L; MEM 4 and 16 mg/L; AMK 8 and 16 mg/L; and LEV 7 and 10 mg/L (12-14). The TGC combinations used were 1 dose of TGC combined with 1 dose of COL, MEM, AMK, or LEV. A 5×105 CFU/mL inoculum of the tested organism was used to inoculate 10 mL of the corresponding broth (containing the antibiotics alone or in combination). The samples were obtained aseptically at predetermined timepoints (0, 2, 4, 8, 12, and 24 hours). Time-kill curves were then constructed as a function of time, and the results are represented as the difference in log10 between the CFU/mL at 0 and 24 hours. A decrease of ≥3 log10 compared with the initial CFU/mL indicated a bactericidal effect. Bacteriostatic activity was defined as a <3 log10 CFU/mL decrease in colony counts. Regrowth was defined as an increase in colony counts from the previous timepoint (15). Synergistic effects were determined by a decrease of ≥2 log10 CFU/mL when comparing the combined antibiotics with the most active drug at that timepoint, while an increase of >2 log10 was considered antagonism. Additivity and indifference were interpreted as any other outcome that did not meet the criteria for either synergism or antagonism (16).
Molecular detection of resistance genes and homology analysis
Polymerase chain reaction (PCR) tests were used to detect the carbapenemase genes (blaKPC, blaNDM, blaVIM-1, blaVIM-2, blaIMP-4, blaIMP-8, blaOXA-23, blaOXA-24, blaOXA-48, blaOXA-51, and blaOXA-58), extended-spectrum beta lactamase (ESBL) genes (blaCTX, blaTEM, blaACC, and blaSHV), colistin resistance gene (blaMCR-1) and quinolone resistance genes (blaqnrA, blaqnrB, blaqnrS, blaqepA, and blaacc(6’)-Ib) in 10 CRKP isolates, as described in a previous study (17). Bioedit software was used to analyze the test data, and the results were compared using online blast software. In addition, multilocus sequence typing (MLST) was performed using 7 housekeeping genes of K. pneumoniae which were amplified using primers Available online online databases (http://bigsdb.pasteur.fr/klebsiella/primers_used.html). The products of PCR were sequenced. Sequence types (STs) were determined using online database tools.
Results
Bacterial isolates
From October 2015 to January 2019 a total of 40 nonduplicated CRKPs were isolated from various clinical specimens. The CRKP isolates were isolated, one from each patient, with an age range of 16–86 years (median 61.5 years). Of the 40 patients, 25 (62.5%) were male. The mortality of the patients with CRKP infections was 20%. Three patients were excluded: 1 outpatient had no hospital records, and 2 nosocomial patients fell out of contact. The specimens with positive culture for CRKP included sputum (80%, n=32), blood (15%, n=6), wound secretion, and other specimens (2.5%, n=1, each). The CRKP isolates emerged from neurosurgery (32.5%, n=13); ICU (50.0%, n=20); emergency room (7.5%, n=3); and hematology, orthopedics, general surgery, and cardiac surgery departments (2.5%, n=1, each).
Antimicrobial susceptibility
In terms of the susceptibility patterns determined by the micro-broth dilution method, the MICs range of CRKP isolates against TGC, COL, MEM, AMK, and LEV were 0.5–2, 2–32, 4–256, 1–16,384, and 0.5–64 µg/mL, respectively. TGC was sensitive against all CRKP isolates. The resistance to COL, MEM, AMK, and LEV was 92.5%, 100%, 2.5%, and 15.0%, respectively. The 50% MIC (MIC50) of TGC, COL, MEM, AMK, and LEV were 1, 8, 16, 4, and 1 µg/mL, respectively. The 90% MIC (MIC90) of TGC, COL, MEM, AMK, and LEV were 2, 16, 128, 8, and 8 µg/mL, respectively (Table 1).

Full table
Synergistic activity and statistical analysis
Synergism and additive effects were detected in TGC combinations using the checkerboard technique, in which the FICI of the TGC-COL, TGC-MEM, TGC-AMK, and TGC-LEV combinations were 0.675±0.188, 0.613±0.358, 0.575±0.237, and 0.863±0.314, respectively (Table 2). TGC-AMK showed the best synergistic and additive effects with a FICI of 70% and 30% in TGC combinations, respectively. The synergistic effects of TGC-MEM and TGC-COL were 50% and 10%, respectively. The additive effects of TGC-COL, TGC-LEV, and TGC-MEM were 80%, 70%, and 40%, respectively. The indifferent effects of TGC-LEV, TGC-COL, and TGC-MEM were 30%, 10%, and 10%, respectively (Table 3). The TGC-COL, TGC-MEM, TGC-AMK, and TGC-LEV combinations decreased the MICs of TGC by 2.7-fold, 4.2-fold, 3.6-fold, and 3.1-fold, respectively.
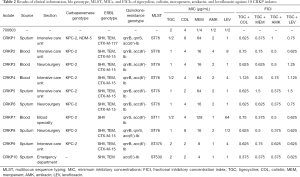
Full table

Full table
Bacterial time-kill effect
Considerable regrowth occurred in the antibiotic monotherapies in the time-kill assay (Figure 1). TGC had bacteriostatic activity and AMK had bactericidal effect on 4 CRKP isolates. On 3 CRKP isolates, 0.25 mg/L COL had bacteriostatic activity, and had a bactericidal effect for 1 CRKP isolate (ST375). On 2 CRKP isolates (ST76 and ST530), 1 mg/L COL had bacteriostatic activity, while on the other 2 CRKP isolates, it had a bactericidal effect. MEM had bacteriostatic activity on 2 CRKP isolates (ST11 and ST530) and had bactericidal effect on the other 2 CRKP isolates. Seven mg/L LEV had bacteriostatic activity on 4 CRKP isolates; 10 mg/L LEV had bacteriostatic activity on 3 CRKP isolates and had a bactericidal effect for 1 CRKP isolate (ST11).
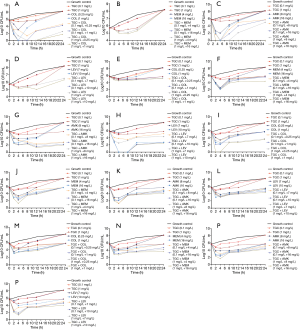
Time-kill assay showed a synergistic effect with TGC combination to 4 CRKP isolates (Figure 1). TGC-COL combination was synergic against 4 CRKP isolates. TGC-MEM combination, TGC-AMK combination, and TGC-LEV combination were synergic against 3 CRKP isolates (ST76, ST375, and ST530), 2 CRKP isolates (ST11 and ST76) and 2 CRKP isolates (ST76 and ST530), respectively.
Molecular detection of resistance genes and homology analysis
Among the 10 CRKP isolates, 9 (90%) were KPC-2 producers. One CRKP isolate (10%) produced NDM-5 carbapenemase gene. ESBL genes were found in 10 (100%), 9 (90%), 7 (70%), and 1 (10%) CRKP isolates carrying blaSHV, blaTEM, blaCTX-M-15, and blaCTX-M-177 genes, respectively. No CRKP isolate was carrying the blaMCR-1 gene. Quinolone resistance genes were found in 10 (100%), 8 (80%), and 2 (20%) CRKP isolates carrying blaacc(6’)-Ib, blaqnrB, and blaqnrS genes, respectively. Four distinct STs were observed among all CRKP isolates and ST76 (n=7) was the most predominant clone, followed by ST11 (n=1), ST375 (n=1), and ST530 (n=1) (Table 2).
Discussion
Despite the increasing occurrence and the severity of infections due to CRKP, limited data exist on the efficacies of the available treatment schemes. We carried out this study to investigate both the in vitro activity of TGC, COL, MEM, AMK, and LEV, alone or in TGC combinations against CRKP, and to provide available treatment schemes applicable to clinical settings.
As for the antimicrobial susceptibility testing in this study, TGC was the most sensitive against all CRKP isolates and was the most effective regimen when used alone. These findings are consistent with the results of another recent survey that included 18 European countries, in which a susceptibility rate of TGC to CRE of 88.6% was found (18). With the investigation of TGC susceptibility in numerous selected pathogens, TGC was found to be one of the most active antimicrobial agents against gram-positive isolates and also to be effective against gram-negative isolates in vitro, including drug-resistant pathogens (19). Other drugs in our study with higher sensitivities were AMK and LEV. AMK had a higher sensitivity rate, possibly because it has only been used for a short time in this region or it has been re-enabled somehow. When AMK is used to treat CRKP, it produces aminoglycoside-modifying enzymes and target 16S rRNA gene mutations cannot be ignored (20). The sensitivity of LEV to CRKP was 85%, and some isolates may be resistant as they carry the quinolone-resistance genes like the blaacc(6’)-Ib, blaqnrB, and blaqnrS genes. However, there were other drugs with higher resistance including COL and MEM. The resistance rate to COL was over 92.5% and was negative for the blaMCR-1 gene. The blaMCR-1 gene had a low level of resistance to COL (8–16 µg/mL), but COL resistance may have the PmrA/PmrB and PhoP/PhoQ two-element regulation systems. All CRKP isolates were resistant to MEM, and most of them carried carbapenemase genes and ESBL genes that might have led to the resistance. CRKP isolates carrying multiple resistance genes render most antibiotic monotherapies ineffective. Meanwhile, it was noted that the higher dose regimen of either antibiotic monotherapy may not be tolerated due to serious side effects, which also becomes an important factor in discussing an optimal combination dosing regimen. Combinations may be the best choice for CRKP treatment through choosing an optimal combination to reduce the standard dose or use a lower dose in combinations.
In this study, we have used 4 antibiotic combinations and found synergism or additive effect for all of these combinations against most of CRKP isolates in the checkerboard microdilution. Overall, the combination of TGC-AMK was the most effective regimen, demonstrating synergism and additive effects against all 10 CRKP isolates. TGC and AMK both work together in bacterial ribosome by binding and interfering with bacterial protein synthesis to kill bacteria effectively (21,22). High-sensitive drugs may have better antibacterial effects in combinations, and combinations can have positive synergism and additive effects on CRKP isolates carrying multi-drug resistance genes. At the same time, our study found that the combination of TGC-COL and TGC-MEM were the best combinations against 4 CRKP isolates in the time-kill assay. The combination of TGC-COL had a synergic effect against KPC-2-producing KP, which is consistent with the result that Toledo et al. reported; it did, however, have an antagonistic effect in combination with TGC-MEM, which is inconsistent with Toledo et al.’s reporting (23). At the same time, the clinical research conducted by Bi et al. showed that a TGC-MEM combination treatment failed in an infection caused by CRKP (24). Therefore, TGC-MEM combinations should be used with caution. The TGC-LEV combination only showed additive effect and even had an indifferent effect in the checkerboard microdilution while showing a synergistic effect against only one CRKP isolate in the time-kill assay. We also found that the 4 combinations had a synergic or additive effect against ST76 CRKP isolates in the checkerboard microdilution and showed a synergic effect in the time-kill assay.
The majority of CRKP isolates produced KPC-2 in our study. First isolate of the KPC-2-producing KP was isolated in the First Affiliated Hospital of Zhejiang University School of Medicine in China in 2004 (25). Carbapenem resistance to KP has increased globally during the past decade and is typically caused by carbapenemase production in particular. KPC-2-producing CRKP isolates are of great clinical concern because of the frequent co-resistance to multiple antibiotic classes. Clinical studies have concluded that combination antibiotic therapy is associated with a better outcome than monotherapy for the treatment of severe infections with these isolates even if the isolated bacteria are susceptible to the individual drugs in vitro (26). This study showed that the 4 combinations had a synergistic or additive effect on most of KPC-2-producing CRKP isolates in the checkerboard microdilution, and the combination of TGC-COL, TGC-MEM, and TGC-AMK had a synergistic effect in the time-kill assay. The part of KPC-2-producing CRKP isolates were indifferent to all combinations except for the TGC-AMK combination in the checkerboard microdilution, indicating that the presence of the KPC gene resulted in a weakened antibacterial effect in combinations, but on 1 CRKP isolate without the KPC-2 gene, the combination showed a synergic or additive effect in the checkerboard microdilution and had a synergistic effect in the time-kill assay. In vitro experiments can reduce the MIC value of drug monotherapy against CRKP isolates and have better antibacterial activity in combinations. Assessment conducted by using the mean value of the steady-state concentrations of the nonprotein-bound drug in humans against CRKP isolates showed that combinations had a synergistic effect. Therefore, the optimal combination for the KPC-2-producing CRKP isolates should be selected and used with caution.
This study also found that the CRKP9 isolate was a hypervirulent Klebsiella pneumoniae (hvKP) while the capsular serotyping was K2. CRKP9-associated virulence genes mainly included the capsular polysaccharide gene rmpA, siderophore-associated genes iucBC, iutA, iroBD, and aerobactin were present on the tig00000014 plasmid; fimbrial adhesin genes, fimA-H and mrkD, and siderophore-associated genes, iutA and entAB, were present on the chromosome. CRKP9 isolate was sensitive to the 5 tested antibiotic monotherapies in the microbroth dilution and not sensitive to MEM. In line with these findings, Liao et al.’s earlier study reported that ST375 belonged to the K2 serotype and was sensitive to most antibiotics (27). TGC combinations had additive effects against the CRKP9 isolate in the checkerboard microdilution, the combinations of TGC-COL and TGC-MEM had synergistic effects in the time-kill assay, and CRKP isolates carrying virulence-associated genes were more sensitive in both monotherapy or combination. However, he CRKP9 isolate, which was highly sensitive to serum complement-mediated killing, died within 3 hours. Whether the addition of serum complement in antibiotics can increase the antibacterial activity against hvKP isolates remains to be studied.
Among the few antibiotics that remain effective, COL, TGC, gentamicin (GEN), and fosfomycin (FM) are often used in combination therapy against CRKP isolates (28). However, the most recommended treatment regimens are confusing, and there is no consensus regarding which antimicrobial combinations should be used to treat these infections (13). In our previously study, COL combined with MEM and AMK showed synergistic and additive effects against CRKP isolates and decreased the MICs of COL by 5.8- and 5.3-fold, respectively. In vitro antibacterial activity of COL combination was better than the TGC combination because of the greater reduction in the MICs of the main antibiotic in combination. However, the combination of TGC with MEM or AMK showed synergistic and additive effects, and even indifferent effects and a decrease of the MICs of TGC by 4.2- and 3.6-fold, respectively. TGC has been approved in recent years for clinical use in China, but resistance to TGC has emerged since its approval (29,30). Du et al. confirmed the transfer capacity of TGC resistance, which was found to be mediated by the mutated tetA through a transferable plasmid. This finding serves as a therapeutic warning, as the tetA gene is frequently carried by CRKP isolates (31). However, a COL combination can reduce the development of COL heteroresistance as proven in in vitro studies, but this has yet to be supported in clinical studies (32). Therefore, COL combination is superior to TGC combination in the treatment of infection caused by CRKP.
Based on our study’s conclusions, we suggest to physicians that TGC combined with AMK is the last active group in the setting of low-dose regimens. Conversely, the combination of TGC with COL or MEM showed the most in vitro synergistic and bactericidal activities against the ESBL-producing, carbapenem-resistant, and even virulence-associated gene CRKP isolates; this regimen could be considered as a last resort approach for the infections caused by CRKP isolates. In addition, TGC combination might circumvent the overuse of carbapenems during the era of multi-drug resistance in KP.
Acknowledgments
We thank the team of curators at the Institute Pasteur MLST system (Paris, France) for importing novel alleles, profiles, and/or isolates on http://bigsdb.web.pasteur.fr.
Funding: This work was supported by Excellent Team of Young Teachers Foundation of Heilongjiang Province (2018-KYYWF-0916), Research Project of Yongchuan Hospital Affiliated to Chongqing Medical University (YJYJ201902), Heilongjiang Provincial Health and Family Planning Commission on Scientific Research Project (2018-290 and 2018-291).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study protocol was approved by the Ethics Committee of Jiamusi University Clinical Medical College for research. The committee’s reference number is 0326. Individual informed consent was waived by the ethics committee listed above because this study used currently existing sample collected during the course of routine medical care and did not pose any additional risks to the patients.
References
- Satlin MJ, Chen L, Patel G, et al. Multicenter Clinical and Molecular Epidemiological Analysis of Bacteremia Due to Carbapenem-Resistant Enterobacteriaceae (CRE) in the CRE Epicenter of the United States. Antimicrob Agents Chemother 2017. [Crossref] [PubMed]
- El-Gamal MI, Brahim I, Hisham N, et al. Recent updates of carbapenem antibiotics. Eur J Med Chem 2017;131:185-95. [Crossref] [PubMed]
- Ni W, Li G, Zhao J, et al. Use of Monte Carlo simulation to evaluate the efficacy of tigecycline and minocycline for the treatment of pneumonia due to carbapenemase-producing Klebsiella pneumoniae. Infect Dis (Lond) 2018;50:507-13. [Crossref] [PubMed]
- Kaye KS, Pogue JM, Tran TB, et al. Agents of Last Resort: Polymyxin Resistance. Infect Dis Clin North Am 2016;30:391-414. [Crossref] [PubMed]
- Hu Y, Liu F, Lin IY, et al. Dissemination of the mcr-1 colistin resistance gene. Lancet Infect Dis 2016;16:146-7. [Crossref] [PubMed]
- Kinch MS, Patridge E, Plummer M, et al. An analysis of FDA-approved drugs for infectious disease: antibacterial agents. Drug Discov Today 2014;19:1283-7. [Crossref] [PubMed]
- Nosengo N. Can you teach old drugs new tricks? Nature 2016;534:314-6. [Crossref] [PubMed]
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; twenty-sixth informational supplement. Document M100-S26. Wayne, PA: CLSI, 2016. Available online: http://www.clsi.org/
- European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 6, 2016. Available online: http://www.eucast.org/
- Bai Y, Liu B, Wang T, et al. In vitro activities of combinations of rifampin with other antimicrobials against multidrug-resistant Acinetobacter baumannii. Antimicrob Agents Chemother 2015;59:1466-71. [Crossref] [PubMed]
- Clinical and Laboratory Standards Institute. Methods for determining bactericidal activity of antimicrobial agents. Document M26-A. Wayne, PA: CLSI, 1999. Available online: http://www.clsi.org/
- Kulengowski B, Rutter WC, Campion JJ, et al. Effect of increasing meropenem MIC on the killing activity of meropenem in combination with amikacin or polymyxin B against MBL and KPC producing Enterobacter cloacae. Diagn Microbiol Infect Dis 2018;92:262-6. [Crossref] [PubMed]
- Yu W, Shen P, Bao Z, et al. In vitro antibacterial activity of fosfomycin combined with other antimicrobials against KPC-producing Klebsiella pneumoniae. Int J Antimicrob Agents 2017;50:237-41. [Crossref] [PubMed]
- Garrison MW. Comparative antimicrobial activity of levofloxacin and ciprofloxacin against Streptococcus pneumoniae. J Antimicrob Chemother 2003;52:503-6. [Crossref] [PubMed]
- Doern CD. When Does 2 Plus 2 Equal 5? A Review of Antimicrobial Synergy Testing. J Clin Microbiol 2014;52:4124-8. [Crossref] [PubMed]
- Petersen PJ, Labthavikul P, Jones CH, et al. In vitro antibacterial activities of tigecycline in combination with other antimicrobial agents determined by chequerboard and time-kill kinetic analysis. J Antimicrob Chemother 2006;57:573-6. [Crossref] [PubMed]
- Gong X, Zhang J, Su S, et al. Molecular characterization and epidemiology of carbapenem non-susceptible Enterobacteriaceae isolated from the Eastern region of Heilongjiang Province, China. BMC Infect Dis 2018;18:417. [Crossref] [PubMed]
- Sader HS, Castanheira M, Flamm RK, et al. Tigecycline activity tested against carbapenem-resistant Enterobacteriaceae from 18 European nations: results from the SENTRY surveillance program (2010–2013). Diagn Microbiol Infect Dis 2015;83:183-6. [Crossref] [PubMed]
- Xie J, Wang T, Sun J, et al. Optimal tigecycline dosage regimen is urgently needed: results from a pharmacokinetic/pharmacodynamic analysis of tigecycline by Monte Carlo simulation. Int J Infect Dis 2014;18:62-7. [Crossref] [PubMed]
- Galimand M, Courvalin P, Lambert T. Plasmid-mediated high-level resistance to aminoglycosides in Enterobacteriaceae due to 16S rRNA methylation. Antimicrob Agents Chemother 2003;47:2565-71. [Crossref] [PubMed]
- Tarchini G. Tigecycline and bacteremia--the dangers of post hoc analysis of pooled data. Clin Infect Dis 2010;51:867-8. [Crossref] [PubMed]
- Wasserman MR, Pulk A, Zhou Z, et al. Chemically related 4,5-linked aminoglycoside antibiotics drive subunit rotation in opposite directions. Nat Commun 2015;6:7896. [Crossref] [PubMed]
- Toledo PV, Aranha AA Junior, Arend LN, et al. Activity of Antimicrobial Combinations against KPC-2-Producing Klebsiella pneumoniae in a Rat Model and Time-Kill Assay. Antimicrob Agents Chemother 2015;59:4301-4. [Crossref] [PubMed]
- Bi S, Yao X, Huang C, et al. Antagonistic effect between tigecycline and meropenem: successful management of KPC-producing Klebsiella pneumoniae infection. Infection 2019;47:497-500. [Crossref] [PubMed]
- Wei ZQ, Du XX, Yu YS, et al. Plasmid-mediated KPC-2 in a Klebsiella pneumoniae isolate from China. Antimicrob Agents Chemother 2007;51:763-5. [Crossref] [PubMed]
- Tängdén T, Hickman RA, Forsberg P, et al. Evaluation of double- and triple-antibiotic combinations for VIM- and NDM-producing Klebsiella pneumoniae by in vitro time-kill experiments. Antimicrob Agents Chemother 2014;58:1757-62. [Crossref] [PubMed]
- Liao CH, Huang YT, Chang CY, et al. Capsular serotypes and multilocus sequence types of bacteremic Klebsiella pneumoniae isolates associated with different types of infections. Eur J Clin Microbiol Infect Dis 2014;33:365-9. [Crossref] [PubMed]
- Morici P, Florio W, Rizzato C, et al. Synergistic activity of synthetic N-terminal peptide of human lactoferrin in combination with various antibiotics against carbapenem-resistant Klebsiella pneumoniae strains. Eur J Clin Microbiol Infect Dis 2017;36:1739-48. [Crossref] [PubMed]
- He F, Fu Y, Chen Q, et al. Tigecycline Susceptibility and the Role of Efflux Pumps in Tigecycline Resistance in KPC-Producing Klebsiella pneumoniae. PLoS One 2015;10:e0119064. [Crossref] [PubMed]
- Hoban DJ, Bouchillon SK, Johnson BM, et al. In vitro activity of tigecycline against 6792 Gram-negative and Gram-positive clinical isolates from the global Tigecycline Evaluation and Surveillance Trial (TEST Program, 2004). Diagn Microbiol Infect Dis 2005;52:215-27. [Crossref] [PubMed]
- Du X, He F, Shi Q, et al. The Rapid Emergence of Tigecycline Resistance in blaKPC-2 Harboring Klebsiella pneumoniae, as Mediated in Vivo by Mutation in tetA During Tigecycline Treatment. Front Microbiol 2018;9:648. [Crossref] [PubMed]
- Ghafur A, Devarajan V, Raja T, et al. Monotherapy versus Combination Therapy against Nonbacteremic Carbapenem-resistant Gram-negative Infections: a Retrospective Observational Study. Indian J Crit Care Med 2017;21:825-9. [Crossref] [PubMed]


